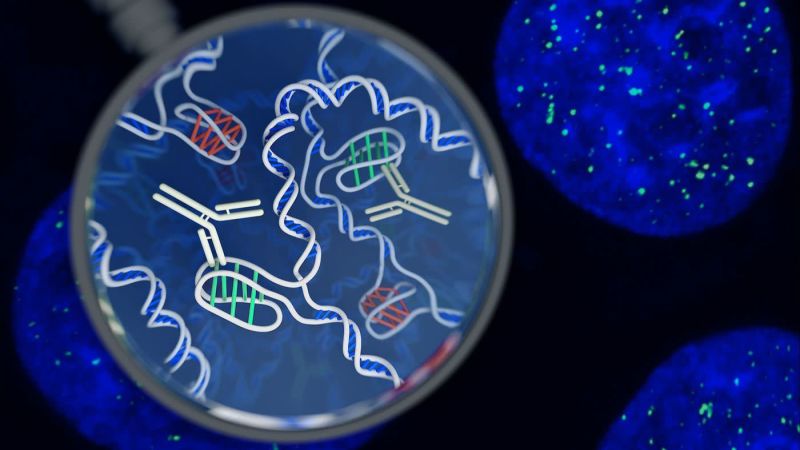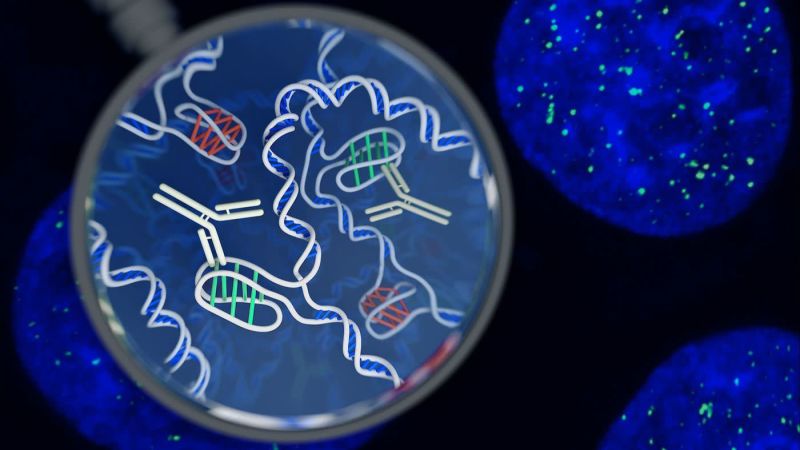
A new structure of DNA has been discovered in living cells for the first time by a team of Australian researchers, opening avenues for unprecedented research into the complex processes that manage the genome’s vast library of information and instructions.
The four-stranded knots – called i-motifs – are a type of secondary shape that were first observed in laboratory samples of DNA in 1993. DNA strands are formed of two long chains of interconnected molecules, or nucleotides, that come in four flavors: adenine (A), guanine (G), thymine (T), and cytosine (C). I-motifs were noted to occur in sequences of the genome where there are many cytosines. Within these stretches, the cytosines deviate from the norm and form bonds with each other instead of linking up with a guanine on the opposite strand.
Yet because these shapes were only seen under artificial, high-pH conditions, i-motifs were initially dismissed as curious chemical rearrangements in response to acidity.
In recent years, however, scientists began to suspect that i-motifs not only occur naturally, but that they may play a role in regulating which genes get expressed. Of course, finding tiny DNA knots amongst the 6 billion-odd nucleotide base pairs in a human cell is no easy task.
The Garvan Institute of Medical Research group’s breakthrough, published in Nature Chemistry, came after they created a sort of molecular bloodhound: An antibody fragment that is perfectly shaped to attach to any i-motifs it encounters inside the nucleus of a cell. Once bound, the scientists could visually confirm the presence of any i-motifs because the antibody is fused to a fluorescent dye.